Difference between revisions of "SMHS LinearModeling LMM Assumptions"
(Created page with "== Linear mixed effects analyses - Mixed Effect Model Assumptions== First review the SMHS_LinearModeling_LMM | Linear mixed effects analyses sec...") |
|||
| Line 43: | Line 43: | ||
# library("ggplot2") | # library("ggplot2") | ||
# ggpairs (hdp[, c("IL6", "CRP", "LengthofStay", "Experience")]) | # ggpairs (hdp[, c("IL6", "CRP", "LengthofStay", "Experience")]) | ||
| + | |||
| + | [[Image:SMHS_LinearModeling_Fig35.png|500px]] | ||
| + | |||
| + | Are there strong linear relations among the continuous variables? Examine CancerStage and LengthofStay closer. The area of bubbles are proportional to the number of observations with the corresponding values. | ||
| + | |||
| + | Violin plots may be used for continuous predictors. We can render all raw data separated by CancerStage. To reduce overlaying, we can add some random noise (along the x axis) or alternatively set the alpha opacity level. | ||
| + | Note that IL6 and CRP have skewed distributions indicating that we use a square root scale on the y axes. The distributions appear normal and symmetric with long right tails, even after square root transformation. | ||
| + | ggplot(hdp, aes(x = CancerStage, y = LengthofStay)) + | ||
| + | stat_sum(aes(size = ..n.., group = 1)) + | ||
| + | scale_size_area(max_size=10) | ||
| + | |||
| + | [[Image:SMHS_LinearModeling_Fig36.png|500px]] | ||
| + | |||
| + | # install.packages("reshape") | ||
| + | # library(reshape) | ||
| + | tmp <- melt(hdp[, c("CancerStage", "IL6", "CRP")], id.vars="CancerStage") | ||
| + | ggplot(tmp, aes(x = CancerStage, y = value)) + | ||
| + | geom_jitter(alpha = .1) + | ||
| + | geom_violin(alpha = .75) + | ||
| + | facet_grid(variable ~ .) + | ||
| + | scale_y_sqrt() | ||
| + | |||
| + | [[Image:SMHS_LinearModeling_Fig37.png|500px]] | ||
| + | |||
| + | The distribution of continuous variables at each level of the binary outcome to provide a better depiction of the change of the binary variables over levels of continuous variables. | ||
| + | tmp <- melt(hdp[, c("remission", "IL6", "CRP", "LengthofStay", "Experience")], id.vars="remission") | ||
| + | ggplot(tmp, aes(factor(remission), y = value, fill=factor(remission)))+ geom_boxplot() + facet_wrap(~variable, scales="free_y") | ||
| + | |||
| + | [[Image:SMHS_LinearModeling_Fig38.png|500px]] | ||
| + | |||
| + | <b>Types of Data Analyses</b> | ||
| + | *Mixed effects logistic regression, the focus of this page. | ||
| + | *Mixed effects probit regression is very similar to mixed effects logistic regression, but it uses the normal CDF instead of the logistic CDF. Both model binary outcomes and can include fixed and random effects. (Note: This link function, aka Probit link, defined in the 1930’s by biologists studying the dosage-cure rate link, refers to the “probability unit”. It’s kind of the inverse CDF, of the model: if Y = Φ(Xβ+ ε), then Probit link = Φ^(-1) (Y). | ||
| + | *Fixed effects logistic regression is limited in this case because it may ignore necessary random effects and/or non-independence in the data. | ||
| + | *Fixed effects probit regression is limited in this case because it may ignore necessary random effects and/or non-independence in the data. | ||
| + | *Logistic regression with clustered standard errors. These can adjust for non-independence but does not allow for random effects. | ||
| + | *Probit regression with clustered standard errors. These can adjust for non-independence but does not allow for random effects. | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
Revision as of 17:27, 17 February 2016
Contents
Linear mixed effects analyses - Mixed Effect Model Assumptions
First review the Linear mixed effects analyses section.
The same conditions we have in the fixed effect multivariate linear model apply to mixed and random effect models – co-linearity, influential data points, homoscedasticity, and lack of normality. These assumptions can be checked by creating residual plots, histogram plots of the residuals or a Q-Q normal probability plots.
The fixed effect independence condition is relaxed in mixed/random effect models as this was the main motivation for mixed models – to resolve dependencies in the data. Mixed effect models still require independence, e.g., when ignoring independent and including just a fixed effect for a variable of interest. For instance, working with a model that does not include a random effect “Player”, then we have multiple Weight responses per Player. This would violate the LME model independence assumption. Careful selection of fixed effects and random effects is necessary to resolve potential dependencies in the data.
The function dfbeta() can’t be used for assessing influential data points in mixed effects linear models the way it can for fixed effect models. To check for influential points in mixed effect models the package influence.ME or a leave-one-out validation can be employed.
For example we can define a vector of size equal to the number of rows in the data. Iterating over each row (i), we estimate a new mixed model excluding the current row index (data[-i,]). The function fixef() extracts the coefficients of interest, which can be adapted to the specific analysis. Running fixef() on the linear model yields the position of the relevant coefficient. For example, position “1” refers to the intercept (which is always the first coefficient mentioned in the coefficient table) and position “2” reflects the effect of “Height” appears second in the list of coefficients.
df <- as.data.frame(data)
all.res=numeric(nrow(df))
for(i in 1:nrow(df))
{ # Generic
# myfullmodel=lmer(response~predictor+ (1+predictor|randomeffect))
# results[i]=fixef(myfullmodel)[parameter position index]
fullmodel=lmer(Weight~Height+ (1+Height|Team), data=data[-i,])
results[i]=fixef(fullmodel)[2]
echo ("Row = ", i)
}
Comments
Fixed effects represent explanatory predictors that are expected to have a systematic and predictable influence on the data (response). Whereas random effects represent covariates expected to have a non-systematic, idiosyncratic, unpredictable, or “random” influence on the response variable. Examples of such random effects in experimental studies include “subject/patient/player/unit” and “Age”, as we generally have no control over idiosyncrasies of individual subjects or their age at time of observation.
Often fixed effects are expected to exhaust the population of interest, or the levels of a factor. In the MLB study the factor “Team” may not exhaust the space as there are other teams/leagues. However, for MLB at a fixed time, the “Team” factor may be fully exhaustive. Same with Height. Random effects represent sub-samples from the population of interest and may not “exhaust” the population as more players or teams could be included in the study. The levels of random factor may only represent a small sub-subset of all levels of the factor.
Hands-on Activity
Use these cancer data (http://www.ats.ucla.edu/stat/data/hdp.csv), representing cancer phenotypes and predictors (e.g., "IL6", "CRP", "LengthofStay", "Experience") and outcome measures (e.g., remission) collected on patients, nested within doctors (DID) and within hospitals (HID). To fit a mixed model (http://www.ats.ucla.edu/stat/r/dae/melogit.htm) and examine remissions as cancer outcomes.
This lung cancer dataset includes a variety of outcomes collected on patients, nested within doctors, who are in turn nested within hospitals. Doctor level variables include experience.
hdp <- read.csv("http://www.ats.ucla.edu/stat/data/hdp.csv")
hdp <- within(hdp, {
Married <- factor(Married, levels = 0:1, labels = c("no", "yes"))
DID <- factor(DID)
HID <- factor(HID)
})
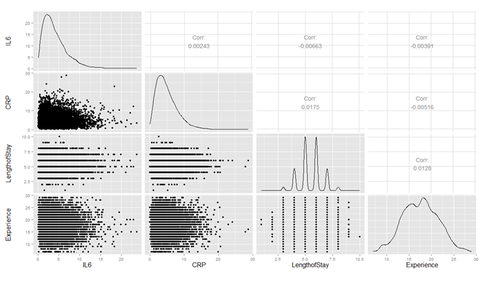
Plot several continuous predictor variables to examine the distributions and catch coding errors (e.g., if values range from 0 to 7, but we see a 999), and explore the relationship among our variables.
# install.packages("ggally")
# library(GGally)
# library("ggplot2")
# ggpairs (hdp[, c("IL6", "CRP", "LengthofStay", "Experience")])
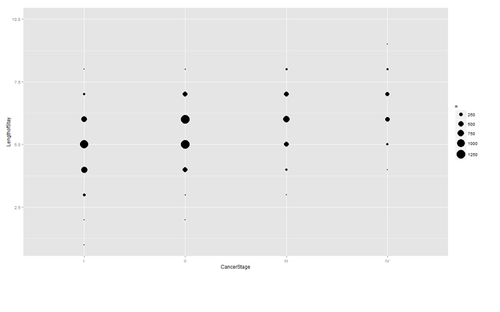
Are there strong linear relations among the continuous variables? Examine CancerStage and LengthofStay closer. The area of bubbles are proportional to the number of observations with the corresponding values.
Violin plots may be used for continuous predictors. We can render all raw data separated by CancerStage. To reduce overlaying, we can add some random noise (along the x axis) or alternatively set the alpha opacity level. Note that IL6 and CRP have skewed distributions indicating that we use a square root scale on the y axes. The distributions appear normal and symmetric with long right tails, even after square root transformation.
ggplot(hdp, aes(x = CancerStage, y = LengthofStay)) + stat_sum(aes(size = ..n.., group = 1)) + scale_size_area(max_size=10)
# install.packages("reshape")
# library(reshape)
tmp <- melt(hdp[, c("CancerStage", "IL6", "CRP")], id.vars="CancerStage")
ggplot(tmp, aes(x = CancerStage, y = value)) +
geom_jitter(alpha = .1) +
geom_violin(alpha = .75) +
facet_grid(variable ~ .) +
scale_y_sqrt()
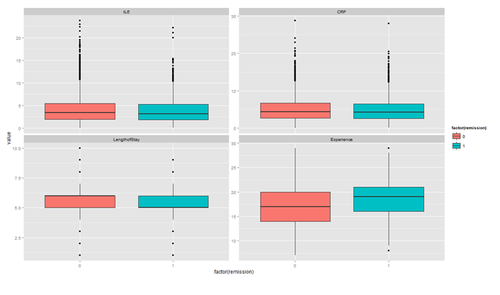
The distribution of continuous variables at each level of the binary outcome to provide a better depiction of the change of the binary variables over levels of continuous variables.
tmp <- melt(hdp[, c("remission", "IL6", "CRP", "LengthofStay", "Experience")], id.vars="remission")
ggplot(tmp, aes(factor(remission), y = value, fill=factor(remission)))+ geom_boxplot() + facet_wrap(~variable, scales="free_y")
Types of Data Analyses
- Mixed effects logistic regression, the focus of this page.
- Mixed effects probit regression is very similar to mixed effects logistic regression, but it uses the normal CDF instead of the logistic CDF. Both model binary outcomes and can include fixed and random effects. (Note: This link function, aka Probit link, defined in the 1930’s by biologists studying the dosage-cure rate link, refers to the “probability unit”. It’s kind of the inverse CDF, of the model: if Y = Φ(Xβ+ ε), then Probit link = Φ^(-1) (Y).
- Fixed effects logistic regression is limited in this case because it may ignore necessary random effects and/or non-independence in the data.
- Fixed effects probit regression is limited in this case because it may ignore necessary random effects and/or non-independence in the data.
- Logistic regression with clustered standard errors. These can adjust for non-independence but does not allow for random effects.
- Probit regression with clustered standard errors. These can adjust for non-independence but does not allow for random effects.
Next See
Machine Learning Algorithms section for data modeling, training , testing, forecasting, prediction, and simulation.
- SOCR Home page: http://www.socr.umich.edu
Translate this page: