SMHS Epidemiology
Contents
Scientific Methods for Health Sciences - Epidemiology
Overview:
After a general introduction to the filed of Epidemiology, students can have a basic idea of the language of Epidemiology. In this course, we want to identify and describe the population patterns of health-related risk factors and health-related outcomes in terms of persons, place and time. We are interested in exploring current major public health issues and try to identify and evaluate the main determinants of such public health issues (e.g. demographic, genetic, infectious, behavioral, and social). With all the concepts and methodologies of analysis in Epidemiology, application would be the next step. Here we examine and apply analytical approaches to data from different epidemiologic study designs (e.g., cross-sectional, cohort, randomized studies) and to critically appraise epidemiological findings.
Motivation:
Goals of this course:
- To understand basic features of the human genome and the distribution of mutations among individuals.
- To understand the principles of segregation and linkage as they apply to human pedigree analysis and the identification of genetic variations associated with disease.
- To learn population and quantitative genetic concepts that are necessary in order to study the relationship between genetic variation and disease variation in populations.
- To learn about prototypical gene-disease relationships that are important to public health.
- To understand the key issues in genetic testing in populations.
- To understand the genetic complexity of common chronic disease.
- To have a basic understanding of the importance and biological basis of epigenetic mechanisms, gene-environment interactions, and gene-gene interactions.
Theory
- Public Heath Genetics:some current and potential applications of genome research include:
- Molecular medicine: improved diagnosis of disease; earlier detection of genetic predispositions to disease; rational drug design; gene therapy and control systems for drugs; pharmacogenomics “custom drugs”.
- Microbial genomics: new energy sources (biofuels); environmental monitoring to detect pollutants; protection from biological and chemical warfare; safe, efficient toxic waste cleanup.
- Risk assessment: assess health damage and risks caused by radiation exposure, include low-dose exposures; assess health damage and risks caused by exposure to mutagenic chemicals and cancer causing toxins.
- Bio-archaeology, anthropology, evolution, and human migration: study evolution through germline mutations in lineages; study migration of different population groups based on X chromosome or Y chromosome; compare breakpoints in the evolution of mutations with ages of populations and historical events.
- DNA forensics (identification): identify potential suspects whose DNA may match evidence left at crime scenes; exonerate persons wrongly accused of crimes; identify crime and catastrophe victims; establish paternity and other family relationships; determine pedigree for seed or livestock breeds.
- Agriculture, livestock breeding, and bioprocessing: more nutritious produce; Biopesticides; healthier, more productive, disease-resistant farm animals; new environmental cleanup uses for plants like tobacco.
*The Human Genome and Mutation
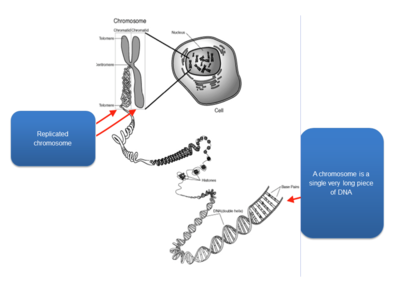
- Chromosomes are highly condensed DNA:
- Chromosomal banding pattern: condensed chromosomes can be stained to create the appearance of dark and light bands; dark bands represent regions rich in As and Ts; each band contains millions of DNA nucleotides; each chromosome has a unique banding pattern.
- A human karyotype: depicts the entire chromosomal constitutions of a person; normal karyotypes have 46 chromosomes; we get 23 chromosomes from each parent (22 autosomes and 1 sex chromosome).
- Chromatin: composed of DNA and proteins that are associated with the chromosomes. (1) Euchromatin: lightly condensed DNA; gene rich, often actively transcribed. (2) Heterochromatin: highly condensed DNA; often composed of repetitive DNA elements; centromeres and telomeres.
- Centromeres: large arrays of repeated DNA sequences; spindle fibers attach during mitosis to separate sister chromatids.
- Telomeres: arrays of repeated DNA sequences that are often thousands of bases in length; a “cap” at the end of chromosome to provide stability; due to the way that chromosomes replicated, telomeres, shorten with each cell division in human somatic cells.
- International system for human cytogenetic nomenclature: short arms of a chromosome are labeled; long arms are labeled; chromosome bands are labeled p11, p12, etc. like a zip code; the terminal ends of the chromosomes are labeled ter; where the arms meet in the middle is the centromere.
- Genes are located on chromosomes: there are 45 bands on chromosome 5; chromosome 5 contains 1633 genes; chromosome 5 ~ 181000000 bases long; genes are referred to by their chromosomal location.
- Chromosomal abnormalities: there are two types of abnormalities that can occur on a chromosomal level in humans: (1) structural abnormalities – missing, extra, or rearranged genetic material on one particular chromosome; (2) numerical abnormalities – deviations in the total number of chromosomes that an individual has.
Changes in chromosome structure.
Deletion: 46, XY, del(6) (p16.3) Terminal deletion with breakpoint at 6p16.3 Duplication: 46, XX, dup(1) (q22q25) Duplication of chromosome 1 region q22 to q25 Insertion: 46, XY, ins(2;5) (p13;q21q31) An insertion of chromosome 5q21-31 into chromosome 2p13 Translocation: 46, XX, t(2;6) (q35;p21.3) A balanced reciprocal translocation with breakpoints in 2q35 and 6p21.3 Inversion: 46, XY, inv(11) (p11p15) An inversion on chromosome 11 with breakpoints at p11 and p15
- Mutations: caused by changes in the DNA sequence; there are many different types of mutations; can happen in somatic cells or during development of gametes.
Types of mutations: (1) Nucleotide substitutions, involve an alteration in the sequence but not the number of nucleotides (DNA bases) in a gene; (2) Insertions & Deletions, involve an alteration in the number of nucleotides in a gene; (3) trinucleotide repeats, involves an alteration in the number of times that a certain sequence of three bases repeats itself; (4) Splice Site Variation, involve an alteration in the non-coding region of a gene, which affects the way that parts of the gene sequence are combined to make RNA. Results of mutations: mutations in exons may result in – misspelling of protein (missense), truncation of the protein (nonsense), no effect; mutations in introns may result in – no effect, altered regulations of gene expression, splice site variation.
- Genes in Population:
- Gene pool: all available genetic variation in a population; all potential mating combinations.
- Basic concepts:
- Alleles: the type of genetic variation seen at a particular location on a chromosome. (1) fictional form: big “A” allele, little “a” allele; (2) base pair form: T allele at basepair 71349562 on chromosome 2, C allele at basepair 71349562 on chromosome 2; (3) codon form: Arginine at codon 124, Glycine at codon 124.
- Genotypes: we inherit one allele from our mother and one allele from our father to form our genotype. Variation in a single gene like AA, Aa or aa.
- Haplotypes: it is the combination of alleles that an individual has at multiple sites along a chromosome.
- Allele frequencies: the prevalence of a particular allele in a given population. Allele frequency = $\frac {Number\, of\, alleles\,}{2*(number\, of\, people\,)}$.
- Genotype frequencies: prevalence of a particular genotype in a given population.
- Haplotype frequencies: frequency that a haplotype occurs in different ethnic groups.
- Hardy-Weinberg disequilibrium: when genotype frequencies in a population differ from what would be predicted based on allele frequencies.
- Hardy-Weinberg Equilibrium (HWE): in a stable population with random mating, allelic frequencies typically predict genotype frequencies using the law of independent segregation. When allele frequencies can accurately be used to predict genotype frequencies in a population, the population is considered to be in a HWE.
- HWE: suppose a SNP that can only be A or C $(p_{A}+p_{C}=1)$, and probability of having A allele is $p_{A}$, C allele is $p_{C}$, then under HWE, the probability of AA genotype = $p_{A}^{2}$, probability of having CC = $p_{C}^{2}$, the probability of having AC = $2p_{A}p_{C}$.
- Steps to test HWE: (1) estimate allele frequencies; (2) calculate the expected relative genotype frequencies under HWE; (3) calculate the expected number of people with each genotype; (4) calculate the difference between observed and expected number of people with each genotype using $χ^2$ formula; (5) sum up the $χ^2$ components and compare the sum to statistical tables to see if there is significant deviation.
| Observed | Expected | $X^{2}$ component | |
| AA | $N_{AA}$ | $p^{2}(N_{Total})$ | $\frac{(O_{AA}- E_{AA})^{2}}{E_{AA}}$ |
| Aa | $N_{AS}$ | $2pq^{2}(N_{Total})$ | $\frac{(O_{Aa}- E_{Aa})^{2}}{E_{Aa}}$ |
| aa | $N_{aa}$ | $q^{2}(N_{Total})$ | $\frac{(O_{aa}- E_{aa})^{2}}{E_{aa}}$ |
| $N_{Total}$ | $N_{Total}$ | Overall $X^{2}$ Statistics |
$N_{AA}$ is the actual number of people in the population with AA genotype, $N_{Total}$ is the number of people in the population, $p^{2}(N_{Total})$ calculated expected number of people with AA genotype. For $χ^{2}$, the null hypothesis $H_{0}$: the population is in HWE. For two alleles, if the overall $χ^{2}$ is less than or equal to 3.84, then p-value is greater than 0.05 and don’t reject $H_{0}$, population is in HWE; if $χ^{2}$ more than 3.84, p-value is less than 0.05, reject $H_{0}$, population is in HWD (disequilibrium).
*Pedigree Analysis and Probability in Genetics
- Mendel’s Law of Segregation: organism carry two copies of each genetic factor; there is segregation of parental factors during gamete formation; each gamete receives one genetic factor from each parent.
- SOCR Home page: http://www.socr.umich.edu
Translate this page: