SMHS MethodsHeterogeneity
Scientific Methods for Health Sciences - Methods for Studying Heterogeneity of Treatment Effects, Case-Studies of Comparative Effectiveness Research
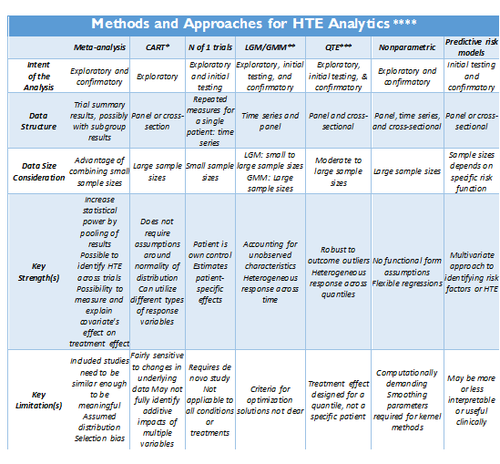
Adopted from: http://dx.doi.org/10.1186/1471-2288-12-185
- *CART: Classification and regression tree (CART) analysis
- ** LGM/GMM: Latent growth modeling/Growth mixture modeling.
- *** QTE: Quantile Treatment Effect.
- **** Standard meta-analysis like fixed and random effect models, and tests of heterogeneity, together with various plots and summaries, can be found in the R-package <bn>rmeta (http://cran.r-project.org/web/packages/rmeta). Non-parametric R approaches are included in the np package, http://cran.r-project.org/web/packages/np/vignettes/np.pdf.
Methods Summaries
Overview
Recursive partitioning is a data mining technique for exploring structure and patterns in complex data. It facilitates the visualization of decision rules for predicting categorical (classification tree) or continuous (regression tree) outcome variables. The R rpart package provides the tools for Classification and Regression Tree (CART) modeling, conditional inference trees, and random forests. Additional resources include an Introduction to Recursive Partitioning Using the RPART Routines . The Appendix includes description of the main CART analysis steps.
install.packages("rpart")
library("rpart")
I. CART (Classification and Regression Tree) is a decision-tree based technique that considers how variation observed in a given response variable (continuous or categorical) can be understood through a systematic deconstruction of the overall study population into subgroups, using explanatory variables of interest. For HTE analysis, CART is best suited for early-stage, exploratory analyses. Its relative simplicity can be powerful in identifying basic relationships between variables of interest, and thus identify potential subgroups for more advanced analyses. The key to CART is its ‘systematic’ approach to the development of the subgroups, which are constructed sequentially through repeated, binary splits of the population of interest, one explanatory variable at a time. In other words, each ‘parent’ group is divided into two ‘child’ groups, with the objective of creating increasingly homogeneous subgroups. The process is repeated and the subgroups are then further split, until no additional variables are available for further subgroup development. The resulting tree structure is oftentimes overgrown, but additional techniques are used to ‘trim’ the tree to a point at which its predictive power is balanced against issues of over-fitting. Because the CART approach does not make assumptions regarding the distribution of the dependent variable, it can be used in situations where other multivariate modeling techniques often used for exploratory predictive risk modeling would not be appropriate – namely in situations where data are not normally distributed.
CART analyses are useful in situations where there is some evidence to suggest that HTE exists, but the subgroups defining the heterogeneous response are not well understood. CART allows for an exploration of response in a myriad of complex subpopulations, and more recently developed ensemble methods (such as Bayesian Additive Regression Trees) allow for more robust analyses through the combination of multiple CART analyses.
Example Fifth Dutch growth study
# Let’s use the Fifth Dutch growth study (2009) fdgs . Is it true that “the world’s tallest nation has stopped growing taller: the height of Dutch children from 1955 to 2009”?
#install.packages("mice")
library("mice")
?fdgs
head(fdgs)
| ID | Reg | Age | Sex | HGT | WGT | HGT.Z | WGT.Z | |
| 1 | 100001 | West | 13.09514 | boy | 175.5 | 75.0 | 1.751 | 2.410 |
| 2 | 100003 | West | 13.81793 | boy | 148.4 | 40.0 | 2.292 | 1.494 |
| 3 | 100004 | West | 13.97125 | boy | 159.9 | 46.5 | 0.743 | 0.783 |
| 4 | 100005 | West | 13.98220 | girl | 159.7 | 46.5 | 0.743 | 0.783 |
| 5 | 100006 | West | 13.52225 | girl | 160.3 | 47.8 | 0.414 | 0.355 |
| 6 | 100018 | East | 10.21492 | boy | 157.8 | 39.7 | 2.025 | 0.823 |
summary(fdgs) summary(fdgs)
| ID | Reg | Age | Sex | HGT |
| Min.:100001 | North:732 | Min.:0.008214 | boy:4829 | Min.:46.0 |
| 1st Qu.:106353 | East:2528 | 1st Qu.:1.618754 | girl:5201 | 1st Qu.:83.8 |
| Median:203855 | South:2931 | Median:8.084873 | Median:131.5 | |
| Mean:180091 | West:2578 | Mean:8.157936 | Mean:123.9 | |
| 3rd Qu.210591 | City:1261 | 3rd Qu.:13.547570 | 3rd Qu.:162.3 | |
| Max:401955 | Max.:21.993155 | Max.:208.0 | ||
| NA's: 23 |
(1) Classification Tree
Let's use the data frame fdgs to predict Region, from Age, Height, and Weight.
# grow tree fit.1 <- rpart(reg ~ age + hgt + wgt, method="class", data= fdgs[,-1])
printcp(fit.1) # display the results plotcp(fit.1) # visualize cross-validation results summary(fit.1) # detailed summary of splits
# plot tree par(oma=c(0,0,2,0)) plot(fit.1, uniform=TRUE, margin=0.3, main="Classification Tree for Region (FDGS Data)") text(fit.1, use.n=TRUE, all=TRUE, cex=1.0)
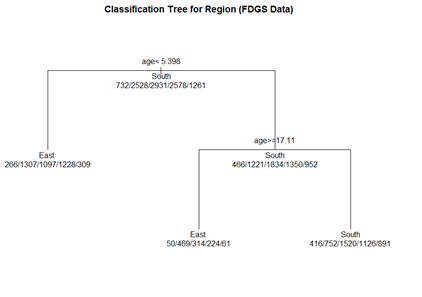
# create a better plot of the classification tree post(fit.1, title = "Classification Tree for Region (FDGS Data)", file = "")
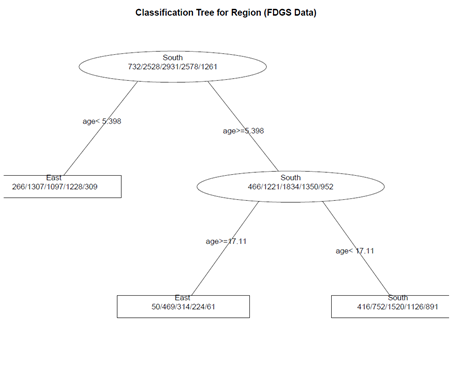
(2) Pruning the tree
pruned.fit.1<- prune(fit.1, cp= fit.1$\$$cptable[which.min(fit.1$\$$\$$cptable[,"xerror"]),"CP"])
# plot the pruned tree
plot(pruned.fit.1, uniform=TRUE, main="Pruned Classification Tree for Region (FDGS Data)")
text(pruned.fit.1, use.n=TRUE, all=TRUE, cex=1.0)
post(pruned.fit.1, title = "Pruned Classification Tree for Region (FDGS Data)")
Not much change, as the initial tree is not complex!
3) Random Forests
Random forests may improve predictive accuracy by generating a large number of bootstrapped trees (based on random samples of variables). It classifies cases using each tree in this new "forest", and decides the final predicted outcome by combining the results across all of the trees (an average in regression, a majority vote in classification). See the <b>randomForest</b> package.
library(randomForest)
fit.2 <- randomForest(reg ~ age + hgt + wgt, method="class", na.action = na.omit, data= fdgs[,-1])
print(fit.2) # view results
importance(fit.2) # importance of each predictor
Note on missing values/incomplete data: If the data have missing values, we have 3 choices:
1. Use a different tool (rpart handles missing values well)
2. Impute the missing values
3. For a small number of missing cases, we can use na.action = na.omit
==='"`UNIQ--h-2--QINU`"'Latent growth and growth mixture modeling (LGM/GMM)===
LGM and GMM represent structural equation modeling techniques that capture inter-individual differences in longitudinal change corresponding to a particular treatment. For instance, patients’ different timing patterns of the treatment effects may represent the underlying sources of HTE. LGM distinguish if (yes/no) and how (fast/slow, temporary/lasting) patients respond to treatment. The heterogeneous individual growth trajectories are estimated from intra-individual changes over time by examining common population parameters, i.e., slopes, intercepts, and error variances. Suppose each individual has unique initial status (intercept) and response rate (slope) during a specific time interval. Then the variances of the individuals’ baseline measures (intercepts) and changes (slopes) in health outcomes will represent the degree of HTE. The LGM-identified HTE of individual growth curves can be attributed to observed predictors, including both fixed and time varying covariates.
LGM assumes that all individuals are from the same population (too restrictive in some cases). If the HTE is due to observed demographic variables, such as age, gender, and marital status, one may utilize multiple-group LGM. Despite its successful applications for modeling longitudinal change, there may be multiple subpopulations with unobserved heterogeneities. Growth mixture modeling (GMM) extends LGM to allow the identification and prediction of unobserved subpopulations in longitudinal data analysis. Each unobserved subpopulation may constitute its own latent class and behave differently than individuals in other latent classes. Within each latent class, there are also different trajectories across individuals; however, different latent classes don’t share common population parameters. Suppose we are interested in studying retirees’ psychological well-being change trajectory when multiple unknown subpopulations exist. We can add another layer (a latent class variable) on the LGM framework so that the unobserved latent classes can be inferred from the data. The covariates in GMM are designed to affect growth factors distinctly across different latent classes. Therefore, there are two types of HTE: 1) the latent class variable in GMM divides individuals into groups with different growth curves; and 2) coefficient estimates vary across latent classes.
<b>Latent variables</b> are not directly observed – they are inferred (via a model) from other actually observed and directly measured variables. Models that explain observed variables in terms of latent variables are called latent variable models. Then the latent (unobserved) variable is discrete, it’s referred to as <b>latent class variable.</b>
Breast Cancer Example: Recall the LMER package, earlier review discussions, where Linear Mixed Model (LMM) are used for longitudinal data to examine change over time of outcomes according relative to predictive covariates. LMM assumptions include:
(i) continuous longitudinal outcome
(ii) Gaussian random-effects and errors
(iii) linearity of the relationships with the outcome
(iv) homogeneous population
(v) missing at random data
The objectives of LGM/GMM models (see <b>Latent Class Mixed Models, lcmm</b> R package) are to extend the linear mixed model estimation to:
(i) heterogeneous populations (relax (iv) above). Use <mark><b>hlme</b> for latent class linear mixed models</mark> (i.e. Gaussian continuous outcome)
(ii) other types of longitudinal outcomes : ordinal, (bounded) quantitative non-Gaussian outcomes (relax (i), (ii), (iii), (iv)). Use <b>lcmm</b> for general latent class mixed models with outcomes of different nature
(iii) joint analysis of a time-to-event (relax (iv), (v)). Use <b>Jointlcmm</b> for joint latent class models with a longitudinal outcome and a right-censored (left-truncated) time-to-event
Let’s use these data (http://www.ats.ucla.edu/stat/data/hdp.csv), representing cancer phenotypes and predictors (e.g., "IL6", "CRP", "LengthofStay", "Experience") and outcome measures (e.g., remission) collected on patients, nested within doctors (DID) and within hospitals (HID).
We can illustrate the latent class linear mixed models implemented in <b>hlme</b> through a study of the quadratic trajectories of the response (remission) with TumorSize, adjusting for CO2*Pain interaction and assuming correlated random-effects for the functions of SmokingHx and Sex. To estimate the corresponding standard linear mixed model using 1 latent class where CO2 interacts with Pain:
# install.packages("lcmm")
library("lcmm")
hdp <- read.csv("http://www.ats.ucla.edu/stat/data/hdp.csv")
hdp <- within(hdp, {
Married <- factor(Married, levels = 0:1, labels = c("no", "yes"))
DID <- factor(DID)
HID <- factor(HID)
})
add a new subject ID column (last column in the data, “ID”), this is necessary for the hmle call
hdp$\$$ID <- seq.int(nrow(hdp))
model.hlme <- hlme(remission ~ IL6 + CRP + LengthofStay + Experience + I(tumorsize^2) + co2*pain + I(tumorsize^2)*pain, random=~ SmokingHx + Sex, subject='ID', data=hdp, ng=1) summary(model.hlme)
Heterogenous linear mixed model fitted by maximum likelihood method hlme(fixed = remission ~ IL6 + CRP + LengthofStay + Experience + I(tumorsize^2) + co2 * pain + I(tumorsize^2) * pain, random = ~SmokingHx + Sex, subject = "ID", ng = 1, data = hdp) Statistical Model: Dataset: hdp Number of subjects: 8525 Number of observations: 8525 Number of latent classes: 1 Number of parameters: 21 Iteration process: Convergence criteria satisfied Number of iterations: 34 Convergence criteria: parameters= 1.2e-09 : likelihood= 8.3e-06 : second derivatives= 2.7e-05 Goodness-of-fit statistics: maximum log-likelihood: -5223.9 AIC: 10489.79 BIC: 10637.86